|
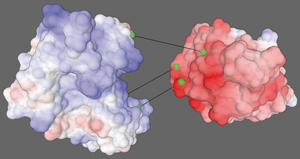
Currently, the most common methods used for docking are fully computational approaches, followed by the use of molecular visualization tools to evaluate results. Studying protein-protein interactions and how proteins form molecular complexes allows researchers to better understand their function in the cell. Protein-Protein docking is a recent practice in biological research which involves using 3D models of proteins to predict the structure of complexes formed by these proteins. The alpha version of Udock is freely accessible at. For most of them, the best scores were obtained with the experimental partner. These favored regions were located inside or nearby the experimental binding interface for 5 out of the 8 proteins in the dataset. The users explored almost all the surface of the proteins that were available in the dataset but favored certain regions that seemed more attractive as potential docking spots. To explore this approach experimentally, we conducted a preliminary two week long playtest where the registered users could perform a cross-docking on a dataset comprising 4 binary protein complexes. We assumed that if given appropriate tools, a naive user's cognitive capabilities could provide relevant data for (1) the prediction of correct interfaces in binary protein complexes and (2) the identification of the experimental partner in interaction among a set of decoys. In Udock, the users tackle simpli fi ed representations of protein structures and explore protein – protein interfaces ’ conformational space using a gami fi ed interactive docking system with on the fl y scoring. Here, we present a new interactive protein docking system, Udock, that makes use of users' cognitive capabilities added up. Protein docking calculations' goal is to predict, given two proteins of known structures, the associate conformation of the corresponding complex. For preconstruction mounting hardware, it is also assumed that drywall is in place and a cutout for the TST‑1080‑DSW has been made.Protein – protein interactions play a crucial role in biological processes. The following procedure assumes that the a TST‑1080‑DSW mounting accessory has been installed completely as described in Install the Mounting Hardware. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
